KAIST
BREAKTHROUGHS
Research Webzine of the KAIST College of Engineering since 2014
Fall 2024 Vol. 23COSINET: Community cohesion looseness in individualized gene networks for precision medicine
Professor Doheon Lee and Dr. Seunghyun Wang developed COSINET (COmmunity COhesion Scores in Individualized gene Network Estimated from single Transcriptomics data), which estimates individualized gene networks from a single gene expression profile and measures community cohesion scores.

As community cohesion plays a critical role in the determination of an individual’s health in social science, could the concept of community cohesion also be applied between genes?
Professor Doheon Lee and Dr. Seunghyun Wang in the Department of Bio and Brain Engineering developed a new technology capable of identifying gene communities which lose their cohesion in individualized gene networks to predict individualized therapeutic targets for precision medicine.
Recently, the incidences of various complex diseases, such as cancer, cardiovascular diseases, and metabolic disorders, have increased significantly due to aging and lifestyle changes. Experts point out that precision medicine, which considers the characteristics of individual patients to enhance treatment efficacy, could be a promising solution to reduce personal and social healthcare costs. To satisfy unmet needs in this area, Professor Doheon Lee's research team has developed COSINET (COmmunity COhesion Scores in Individualized gene Network Estimated from single Transcriptomics data), a technology that precisely estimates individualized gene networks and accurately measures community cohesion scores.
The researchers initially constructed a normal tissue gene network (tissue-specific weighted co-expression network) and found co-expressed communities using hundreds of publicly available normal gene expression profiles. Then, they modeled the correlations of each gene interaction with simple linear regression models and control samples. Next, they estimated the individualized gene network by identifying individualized perturbed interactions that deviated from the co-expression patterns in normal tissues based on the results of the simple linear regression models. Additionally, in each individualized gene network, the community cohesion scores, which precisely quantify the community ability to retain the normal interactions between the genes and their cellular functions, were measured.
The researchers demonstrated that the community cohesion scores can be successfully used to predict the individualized therapeutic targets. First, they prioritized them by finding druggable genes present in the community of which the scores are relatively low in each individual compared to other patients. Then, they pinpointed more individualized therapeutic targets by discovering the genes which greatly contribute to the degree of community cohesion looseness in each individualized gene network.
Professor Doheon Lee stated, "Complex diseases involving multiple genes should be viewed from a systems perspective that considers the interactions between genes rather than focusing on individual genes. The current single-gene-based biomarkers used in clinical practices for precision medicine show limitations in capturing the heterogeneity and complexity of complex diseases. Therefore, the COSINET technology developed in this study, based on the community cohesion looseness in individualized gene networks, could offer a new perspective for precision medicine for complex diseases."
This study, supported by the Ministry of Science and ICT, was published in 'Briefings in Bioinformatics' (May 2024 issue, Oxford University) and was released online on April 15th of that year.
Most Popular

High-performance and sustainable paper coating material that prevents microplastics
Read more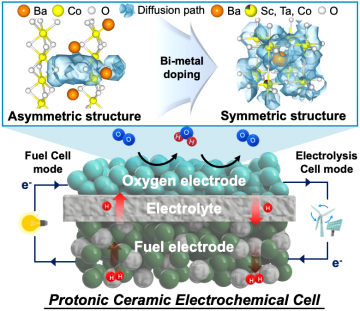
Next-generation ceramic electrochemical cells for global net-zero goals
Read more
Impacts of new town developments on carbon sinks in the Seoul metropolitan area
Read more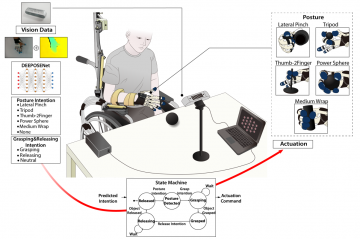
Multi-hand posture rehabilitation for stroke survivors: Rehabilitation system using vision-based intention recognition and a soft robotic glove
Read more
Revolutionizing strength: Hercules artificial muscles 17 times stronger than human muscles
Read more